Dr. Mahlon Collins leads neurodegenerative disease research at the Parallel Squared Technology Institute (PTI). A neurobiologist by training, he has witnessed firsthand the tremendous burden these diseases impose on patients and their families.
During his graduate studies, he worked directly with amyotrophic lateral sclerosis (ALS) patients and their families, collecting cerebrospinal fluid (CSF) samples for biomarker analysis. This required participants undergoing a lumbar puncture, or spinal tap. The process can be difficult: needle insertion is painful and patients often experience headaches afterwards, especially when collecting the large volumes of CSF required for Mahlon’s work.
“Despite this, our patients and their family members enthusiastically donated, often multiple times,” Mahlon remembers. “Their dedication, perseverance, and gratitude for our efforts remains a source of inspiration.” The commitment of patients and their families to do anything they could to advance science continues to drive Mahlon’s work at the intersection of cutting-edge technology and urgent human need.
For Mahlon, the power of proteomics to advance neurodegenerative disease research was immediately apparent. “Simultaneously measuring thousands of proteins immediately struck me as a powerful approach for understanding biological systems,” recalls Mahlon. It was the most exciting technique he encountered as a molecular biology student, and that early fascination has shaped a career dedicated to unravelling the molecular mysteries of diseases like Alzheimer’s and Parkinson’s. “I use proteomics to try to improve our understanding of these diseases, how we diagnose them, and how we develop therapies to treat and prevent them.”
From biomarkers to genetics and back again
Mahlon’s journey through proteomics has been anything but linear. During his PhD, he used mass spectrometry to improve how we diagnose ALS, analysing those hard-won CSF samples with machine learning to identify new ALS biomarkers. But the work also provided other insights into ALS. In particular, the CSF proteome reflected ongoing pathological changes in ALS-afflicted neurons, pointing to a previously unknown form of RNA binding protein pathology in both ALS and Alzheimer’s disease.
Yet the extent of this new pathology, like other ALS pathologies, was highly heterogeneous from patient to patient. For Mahlon, this raised questions about why patients with the same diagnosis can be so pathologically distinct. In part, the answers, Mahlon realised, lay in genetics. But there was a practical problem: mapping genetic influences on protein-related phenotypes would require profiling large samples of genetically distinct individuals. “At the time, this wasn’t practical with mass spec proteomics for most labs,” Mahlon explains, though he notes that researchers like Leonid Kruglyak and Yoav Gilad managed to make it work, producing highly influential findings.
“Lacking their resources and seeing how fluorescent proteins were productively being used to measure phenotypes in thousands of live cells, I made the jump to genomics,” he says. His postdoctoral research explored how individual genetic differences influence molecular processes like protein turnover, and he developed assays to measure genome-wide genetic influences on protein degradation.
“Working on the genetics of protein degradation was a wonderful introduction to the complex ways individual genetic differences shape phenotypic outcomes,” Mahlon explains. His approach revealed something remarkable: genetic variants affect protein degradation pathways in highly substrate-specific ways. “Their magnitude and even direction of effect often depends on which protein is being degraded,” he notes.
Recent advances in mass spectrometry throughput and sensitivity (approaches like plexDIA and PSMtags) have finally made population-scale proteomic studies practical, bringing Mahlon full circle. “There remains much to be discovered about how DNA sequence differences shape molecular processes relevant to human health and disease, including protein degradation, and high-throughput proteomics is poised to deliver important insights in this area,” he says. Now, he’s leading PTI’s efforts to profile the proteomes of individual neurons isolated from human brain tissue.
Breaking through
The technical difficulties of isolating individual neurons from human brain tissue whilst preserving them well enough for proteomic analysis demanded months of persistence, focus, and troubleshooting. Each failed experiment carried extra weight; the tissue came from human Alzheimer’s patients whose commitment to research extended beyond their lifetimes – their final act of donation.
“Every failed one was crushing, especially since we were working with human samples, entrusted to us by patients and their families” Mahlon recalls. The team tried to learn principles using other sample types and sources, hoping to develop their approach more efficiently, but those results didn’t generalise to human post-mortem samples. “It really took a lot of careful refinement and thorough analysis of results from a limited number of samples to get us where we needed to be.”
The technical foundation proved crucial. Dr. Jason Derks, one of PTI’s senior scientists who runs many of their samples, first encountered IonOpticks columns whilst working in Professor Slavov’s lab on single-cell samples. Since then, they’ve become integral to the team’s workflow. “IonOpticks columns have consistently performed well and offered a high degree of robustness and reproducibility, which has served as a reliable foundation for enabling my work,” Derks notes. The columns have proven versatile too, handling everything from bulk samples up to 1 μg down to single organelles like nuclei at just 50 pg. That robustness and reproducibility became even more valuable when working with irreplaceable patient samples.
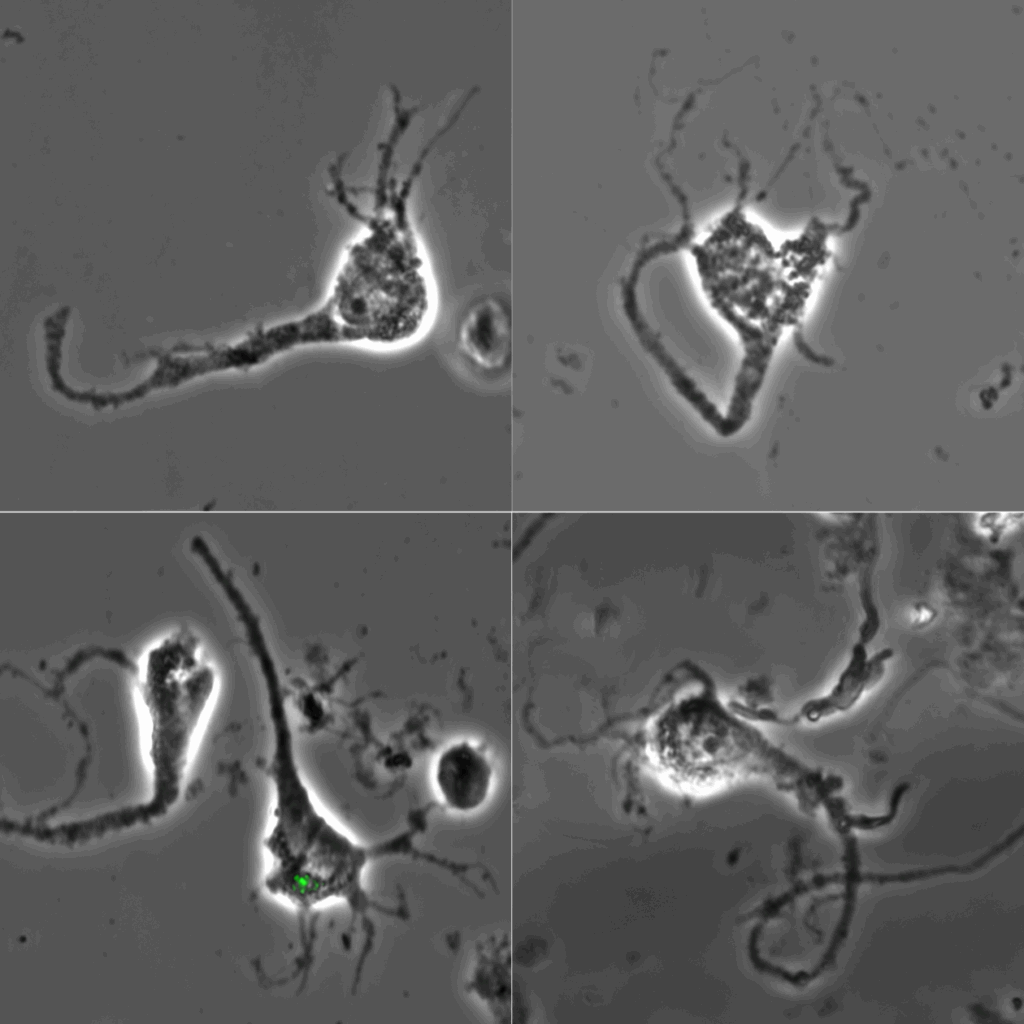
When the breakthrough finally came, when they successfully isolated morphologically identifiable neurons that could be proteomically profiled, the moment was unforgettable. “Seeing those neurons under the microscope for the first time and later looking at their proteomes was and remains deeply exciting and rewarding.” Two things made it exciting, Mahlon explains: “Finally breaking through a challenge that felt insurmountable for months and realising all the exciting things we could now do.”
Those neurons under the microscope represented a view that could unlock new understanding of how neurodegeneration unfolds at the most fundamental level. “Of course, much of that remains to be done,” Mahlon adds, “but we’re excited for what the future holds.”
New insights into Alzheimer’s Disease
That technical capability has already supported significant discoveries. Using plexDIA, Mahlon and colleagues measured proteomic changes in Alzheimer’s disease and uncovered something unexpected: AD results in decreased levels of the 20S proteasome, the catalytic core of the primary eukaryotic protein degradation pathway.
The finding was striking not just for what decreased, but for what increased as a result. The proteins that accumulated in AD “have highly specific subcellular localisations and fast degradation rates, they contain signal sequences that allow them to be targeted for proteasomal degradation, and they are targeted by quality control pathways that recognise mislocalised proteins,” Mahlon explains. The pattern suggests that when the proteasome falters, proteins that depend most heavily on this degradation machinery, particularly those that need to be rapidly cleared or are in the wrong cellular location, begin to accumulate.
The work exemplifies the collaborative strength of the PTI team. “I’m proud of how we leveraged our team’s deep technical expertise in mass spectrometry, proteomics, and computational analysis along with our knowledge of the biology of AD,” Mahlon reflects. “We produced a work that adds new insights into a devastating disease and is technically sound and analytically rigorous, a satisfying achievement.”
Read the full paper here.
The next frontier
Looking ahead, Mahlon sees tremendous potential in PTI’s development of the PSMtags reagent and JMod software. “I’m admittedly biased,” he admits, “but [these tools are] what I’m most excited about in proteomics.” The technology promises to unlock new experimental designs and applications, but Mahlon is particularly enthusiastic about two capabilities: population-scale throughput and enhanced sensitivity for single-cell applications.
For Mahlon, population scale addresses the critical challenges he’s pursued throughout his career. “We use population scale to mean projects that could analyse large samples – such as those from the 1000 genomes project or large cohorts that capture the pathological heterogeneity intrinsic to diseases like ALS and AD. It could also comprise large samples of single cells, such as those that comprise a tissue,” he explains.
These capabilities are essential for uncovering the genetic basis of human health and disease, characterising the extensive patient-to-patient heterogeneity intrinsic to most human diseases, and understanding how complex interactions of multiple cell types and states contribute to a tissue’s normal functions and its dysregulation in disease. For someone who started his career captivated by the idea of measuring thousands of proteins simultaneously, who had to temporarily abandon mass spectrometry because it couldn’t handle the sample sizes genetics demands, and who has dedicated years to understanding patient-to-patient variation, the ability to bring proteomic analysis to population-scale studies represents the convergence of method and mission.
Combined with the sensitivity needed to profile individual cells – those hard-won neurons that took months of failures to successfully isolate – these advances could finally reveal how genetic diversity, cellular heterogeneity, and protein dynamics interweave to create the complex landscape of neurodegenerative disease. By measuring the proteome, now at resolutions from individual neurons to entire populations, made possible by the very samples patients donated, we might be able to understand biological systems well enough to ease the burden of devastating diseases.
Would you like to be featured in an upcoming Community Newsletter? Email us at communications@ionoptickscopy.com and tell us a bit about yourself and your research.