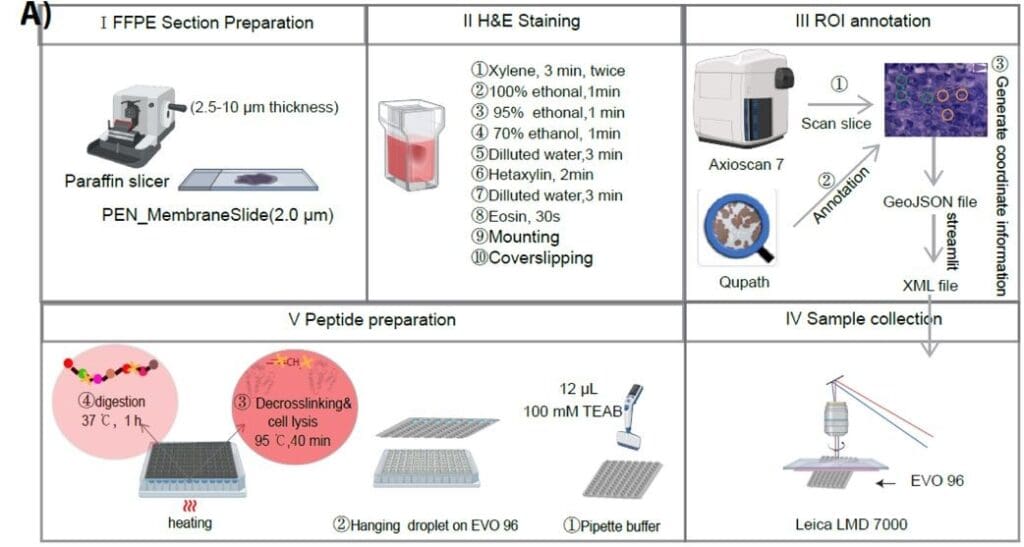
Schematic overview of the high-throughput hanging droplet workflow for micro-FFPE spatial proteomics. From Chen et al., 2025. “High-Throughput Spatial Proteomics at Cellular and Subcellular Resolution via a Hanging Droplet Workflow“, bioRxiv (2025): 2025-11. https://doi.org/10.1038/s41587-025-02791-w; Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Spatial proteomics has emerged as a powerful approach for mapping protein expression within intact tissue architecture, enabling researchers to understand cell-cell interactions and microenvironmental influences that are lost in bulk or single-cell analyses. A major barrier to widespread adoption has been the technical challenge of processing microscale tissue samples from formalin-fixed paraffin-embedded (FFPE) archives. During high-temperature decrosslinking, which is essential for reversing formaldehyde-induced protein crosslinks, tiny samples are prone to rapid evaporation, leading to sample loss, incomplete digestion, and poor reproducibility. Chen et al. aimed to develop a high-throughput, reproducible workflow for spatial proteomics that overcomes evaporation challenges while maintaining sensitivity at cellular and subcellular resolutions.
The researchers from the Olsen lab developed a hanging droplet workflow using the EVO96 proteoCHIP chip, which suspends reaction droplets face-down during heating to prevent evaporation. Laser-capture microdissected FFPE breast cancer samples were processed in parallel (up to 96 samples in approximately 2 hours). The workflow utilised an Evosep One with the 80 SPD Whisper Zoom method (5.6-minute gradient), coupled to an Orbitrap Astral via an EASY-Spray ion source. Separation was performed using an IonOpticks Aurora® Elite™ 15×75 XT C18 UHPLC column in narrow-window data-independent acquisition mode.
This exploratory workflow identified approximately 5,500 protein groups from regions containing ~250 cells, 2,500 protein groups from ~5 cells, and 1,500 protein groups from subcellular structures as small as 60 µm². Applying this method to breast cancer subtypes, the researchers revealed distinct tumour-stromal proteomic programmes across triple-negative, HER2-positive, and luminal subtypes. Tumour regions showed enrichment in RNA processing and proliferative signalling, whilst stromal regions exhibited extracellular matrix remodelling and immune activation signatures.
This scalable approach enables ultrasensitive spatial proteomics from archived FFPE tissues without specialised instrumentation, advancing precision pathology and translational research applications.
Authors
Hao Chen, Ulises H. Guzman, & Jesper V. Olsen;
Title

