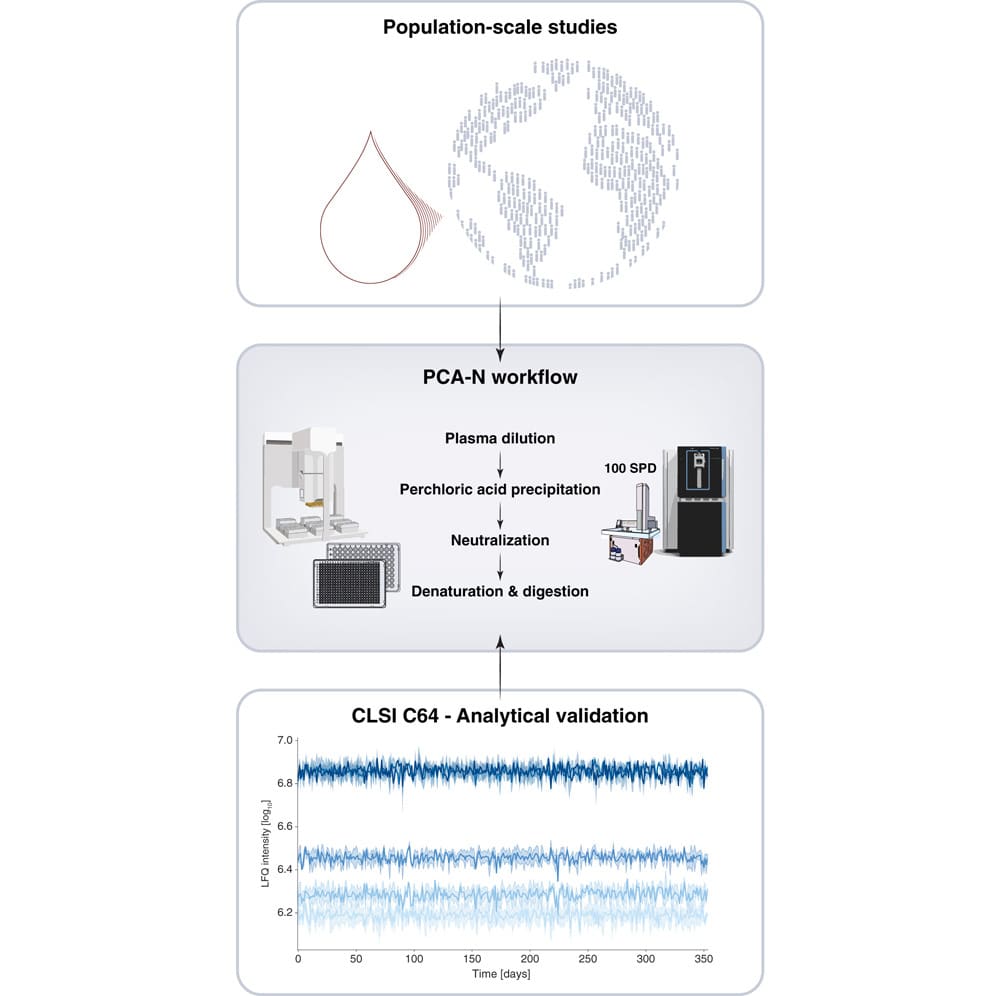
Graphical abstract. From Albrecht et al., 2025. “A Simplified Perchloric Acid Workflow With Neutralization (PCA N) for Democratizing Deep Plasma Proteomics at Population Scale“, Molecular & Cellular Proteomics, Volume 24, Issue 11, 101071; doi: 10.1016/j.mcpro.2025.101071. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Large-scale plasma proteomics studies have been limited by a fundamental trade-off between analytical depth and practical feasibility. While standard undepleted (NEAT) plasma analysis offers simplicity and high throughput, it lacks sufficient depth to detect clinically relevant low-abundance proteins. Alternative techniques introduce various limitations in sample volume requirements, costs, and reproducibility.
To address these challenges, Albrecht et al. developed PCA-N, a novel perchloric acid-based workflow with a neutralisation step that eliminates the need for protein extraction. This innovation drastically reduces sample volume requirements to just 5 µL of plasma while enabling parallelisation to 384-well plates. The streamlined protocol allows preparation of more than 10,000 samples per day at costs similar to standard NEAT plasma analysis.
The researchers from the Mann lab validated PCA-N using an Evosep One coupled to an Orbitrap Astral operated in Data Independent Acquisition (DIA) mode. Peptides were separated on an Aurora Rapid 8×150 XT C18 UHPLC column using an EASY-Spray Source. The quantitative study demonstrated that PCA‑N substantially increases proteome coverage compared to NEAT workflows (approximately 2,000 proteins per sample, with cumulative identifications exceeding 4,000 for larger cohorts), while maintaining comparable reagent costs under 1 USD per sample
The researchers successfully applied the PCA‑N workflow to analyze over 36,000 plasma samples across 311 days, including 1,500 quality control samples interspersed throughout the study, demonstrating strong reproducibility and validating its suitability for extreme‑scale clinical applications. This establishes a new paradigm for population-level proteomics research that could significantly advance biomarker discovery by making deep plasma proteomics accessible and economically feasible for large-scale clinical applications.
Publication
Molecular & Cellular Proteomics (MCP)
Authors
Vincent Albrecht, Johannes B. Müller-Reif, Vincenth Brennsteiner, & Matthias Mann;
Title

