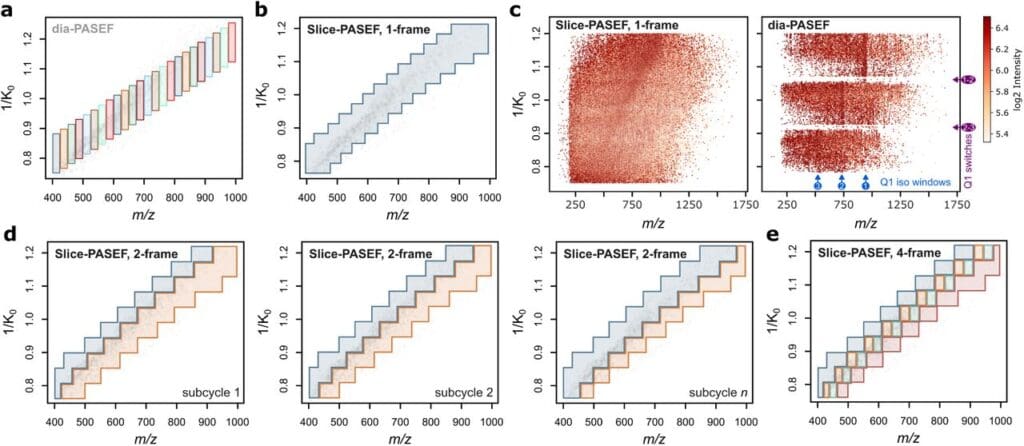
Visualisation of Slice-PASEF methods. From Sinn et al., 2025. “Slice-PASEF: Maximising Ion Utilisation in LC-MS Proteomics“, bioRxiv (2025): 2025-09.; doi: https://doi.org/10.1101/2022.10.31.514544. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Data-independent acquisition (DIA) mass spectrometry has become essential for modern proteomics due to its reproducibility, extensive proteomic coverage and excellent and quantitative performance. However, current methods compromise sensitivity when analysing low sample amounts such as single cells or rare post-translational modifications. To address these limitations and enable deeper proteome coverage from limited material, Sinn et al. developed Slice-PASEF, a novel DIA acquisition strategy that maximizes ion utilization by splitting the precursor ion space into quasi-continuous ‘ion mobility slices’ on timsTOF mass spectrometers. This approach achieves up to a 100% MS/MS duty cycle, compared to the typical 12.5% achieved with conventional dia-PASEF.
To benchmark the Slice-PASEF method, the researchers combined Slice-PASEF with plexDIA for the analyses of U-937 and Jurkat samples using a Vanquish Neo with IonOpticks Aurora Ultimate™ 25 cm x 75 μm CSI C18 UHPLC column, coupled to a timsTOF SCP with CaptiveSpray ion source.
The study demonstrated substantial improvements in proteomic depth: Slice-PASEF identified 84% more precursors and 46% more protein groups from 10 ng samples compared to conventional dia-PASEF. For single-cell proteomics, the method quantified 50% more proteins per cell with 7-fold higher fragment ion signals and improved quantitative precision. In ubiquitinomics applications, Slice-PASEF identified 46% more ubiquitinated peptides from low-input samples.
The enhanced sensitivity enabled comprehensive single-cell proteome profiling with throughput up to 135 cells per day, revealing cell-cycle-specific proteome variations and enabling precise drug mechanism studies. This advancement significantly expands the scope of sensitive proteomics applications, particularly for single-cell analysis and rare post-translational modification studies.
Publication
bioRxiv
Authors
Ludwig R. Sinn, Lukasz Szyrwiel, Justus Grossmann, Kate Lau, Katharina Faisst, Di Qin, Florian Mutschler, Luke Khoury, Andrew Leduc, Markus Ralser, Fabian Coscia, Matthias Selbach, Nikolai Slavov, Nagarjuna Nagaraj, Martin Steger, & Vadim Demichev;
Title

