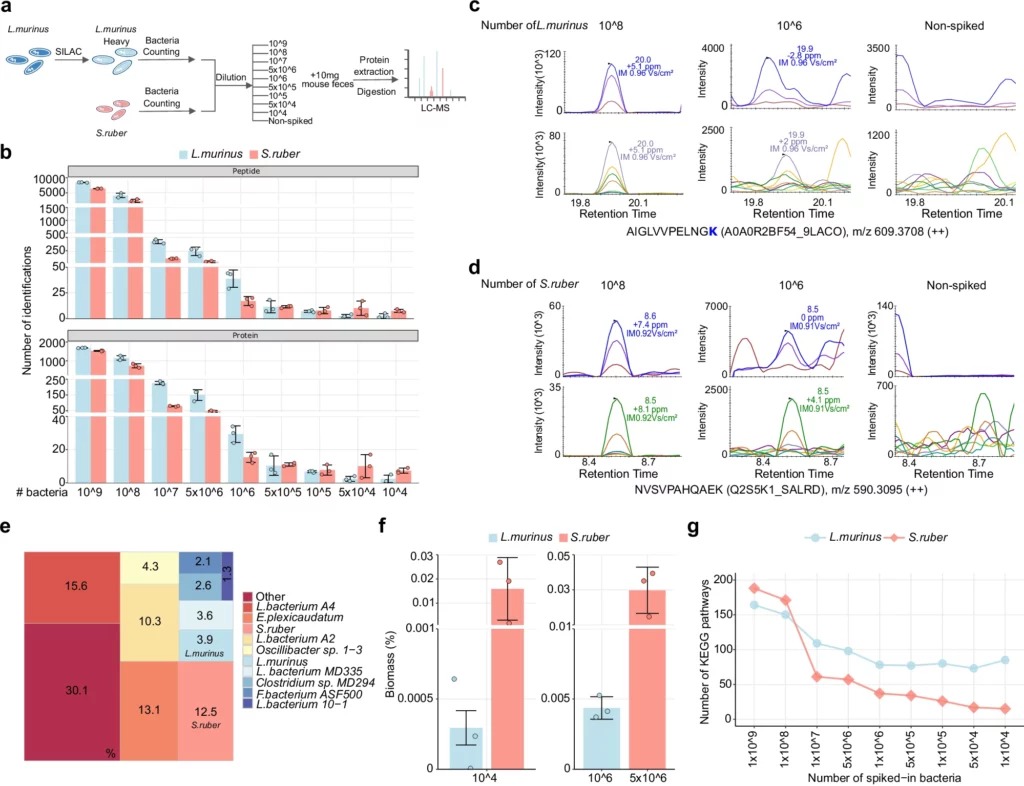
Improving the detection limit of the dark metaproteome using uMetaP workflow. From Xian, F., Brenek, M., Krisp, C. et al. “Ultra-sensitive metaproteomics redefines the dark metaproteome, uncovering host-microbiome interactions and drug targets in intestinal diseases.“, Nat Commun 16, 6644 (2025). https://doi.org/10.1038/s41467-025-61977-7. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Recent work published by Xian et al. in Nature Communications from the Gómez-Varela lab demonstrates the development of uMetaP, an ultra-sensitive metaproteomic solution that leverages cutting-edge technologies to achieve unprecedented sensitivity in exploring complex microbial communities.
Using the mouse gut microbiome as a model, uMetaP was able to quantify 47,925 protein groups and 223 KEGG pathways for both the host and its microbiome. uMetaP detected 220 microbial species, with excellent reproducibility and quantitative precision using low sample amounts (25ng of peptides). As a demonstration of the power of this new technique, this number of identified species is comparable to the average reached by full-length 16S rRNA. These remarkable metrics were achieved by combining uMetaP with the Aurora Ultimate™ 25×75 CSI C18 UHPLC column and timsTOF Ultra.
Through an experimental design that utilises SILAC-labelled bacteria, the researchers addressed two enduring challenges in metaproteomics: the true limit of detection for the dark metaproteome and achieving single-bacterium resolution. Among a theoretical amount of 10 billion spiked-in bacteria, they confidently detected 10,000 spiked-in bacteria. This represents a 5,000-fold reduction in the limit of detection. Moreover, species-specific peptides from a 500-femtogram two-bacteria peptide mix were identified with extraordinary quantification precision and accuracy.
This study highlights the utility of using uMetaP for a broad range of applications including biotech, therapeutic discovery and identification of novel biology, with depth of coverage, sensitivity and reproducibility that allows for complementarity with genomic approaches for investigating microbial ecosystems.
Publication
Nature Communications
Authors
Feng Xian, Malena Brenek, Christoph Krisp, Elisabeth Urbauer, Ranjith Kumar Ravi Kumar, Doriane Aguanno, Tharan Srikumar, Qixin Liu, Allison M. Barry, Bin Ma, Jonathan Krieger, Dirk Haller, Manuela Schmidt & David Gómez-Varela;
Title



