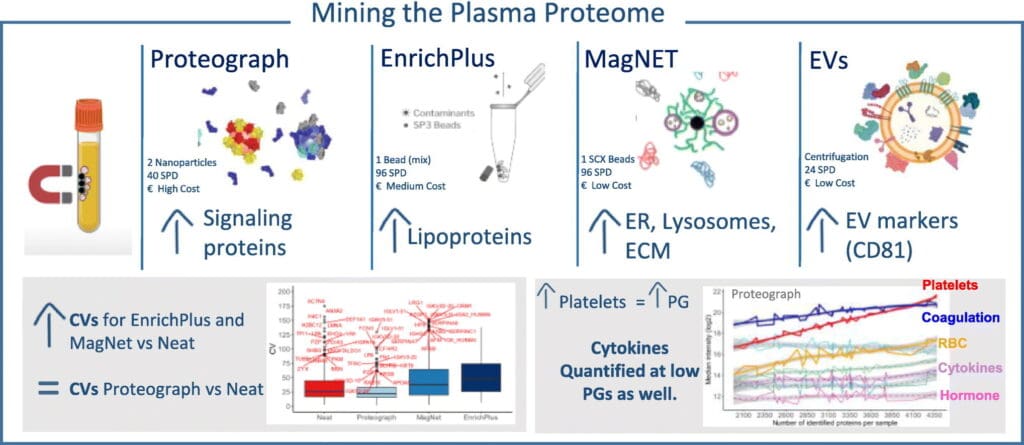
Graphical abstract. From Roger et al., 2025. “Mining the plasma proteome: Evaluation of enrichment methods for depth and reproducibility“, Journal of Proteomics (2025): 105519.; doi: https://doi.org/10.1016/j.jprot.2025.105519. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Deep plasma proteomics faces significant challenges due to the broad dynamic range of protein concentrations, limiting detection of low-abundance biomarkers including extracellular vesicle (EV) cargo proteins, signalling molecules, and tissue leakage proteins. Roger et al. aimed to benchmark three commercial corona formation enrichment strategies – Proteograph Assay XT (Seer), SCX beads (Mag-Net, ResynBio), and functionalised microbeads (ENRICHplus, PreOmics) – against neat plasma and EV fractions to evaluate protein identification depth, quantification reproducibility, and method-specific enrichment patterns.
For plasma set A, an Evosep One was coupled to a timsTOF HT with peptides separated on an Aurora Elite™ 15×75 CSI C18 UHPLC column with the Whisper Zoom 40 SPD method. For plasma set B, samples were analysed on a nanoElute and timsTOF Pro, using an Aurora Ultimate™ 25×75 CSI C18 UHPLC column.
This quantitative proteomics study from the Guerrera lab demonstrated that Proteograph achieved the highest protein identifications with 6.3-fold improvement over neat plasma, whilst maintaining superior quantification reproducibility (median CV 22% vs 38-48% for other methods). The researchers revealed that corona formation strategies reshape the plasma proteome intensity landscape through dynamic range compression, with each method showing distinct protein class preferences—ENRICHplus for lipoproteins, Mag-Net for lysosomal proteins, and Proteograph for cytokines and hormones.
The study established that deeper proteome coverage correlates with platelet contamination levels, yet low-abundance soluble proteins remain detectable regardless of background contamination. This insight guides optimal enrichment strategy selection for biomarker discovery workflows and highlights the importance of evaluating plasma proteomics quality based on cytokine detection rather than total protein count.
Publication
Journal of Proteomics
Authors
K. Roger, I. Metatla, S. Ceccacci, K. Wahbi, L. Motté, C. Chhuon, & I.C. Guerrera;
Title
Mining the plasma proteome: Evaluation of enrichment methods for depth and reproducibility



