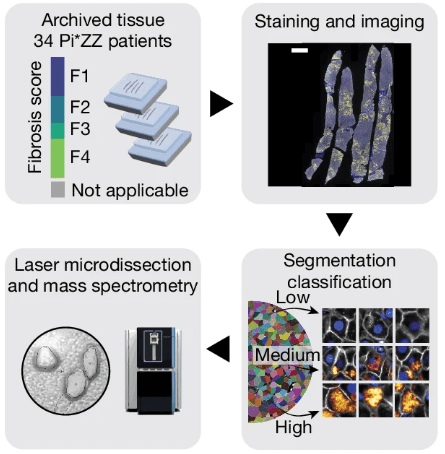
Overview of the Deep Visual Proteomics workflow. Modified from Rosenberger et al., 2025. “Deep Visual Proteomics maps proteotoxicity in a genetic liver disease“, Nature (2025). https://doi.org/10.1038/s41586-025-08885-4. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Alpha-1 antitrypsin deficiency (AATD) is a genetic disorder characterised by protein misfolding and accumulation in hepatocytes, leading to liver fibrosis. Despite having the same disease-causing mutation, patients show diverse clinical outcomes, suggesting unexplored molecular resilience mechanisms. Rosenberger et al. aimed to elucidate the progressive molecular events during proteotoxicity in patient tissues.
The researchers implemented Deep Visual Proteomics (DVP), an innovative approach integrating staining, AI-guided cell segmentation, laser microdissection, and high-sensitivity mass spectrometry. They analysed formalin-fixed paraffin-embedded (FFPE) liver biopsies and resections from patients with the Pi*ZZ genotype across all fibrosis stages.
For mass spectrometry analysis, they used an Evosep One coupled to an Orbitrap Astral equipped with a FAIMS Pro interface and an EASY-Spray source. Peptides were separated on an IonOpticks Aurora Elite XT 15×75 C18 UHPLC column.
This quantitative proteomics study identified up to 5,000 proteins per sample in bulk analyses and up to 4,299 proteins in single-cell analyses. The researchers categorised proteins into early and late responders to proteotoxic stress, revealing a previously unrecognised early peroxisomal response that was significantly delayed in high-grade fibrotic samples.
By integrating morphological information with proteomic data, they identified distinct aggregate morphologies corresponding to different cellular states, with globular aggregates marking late-stage apoptotic cells.
These findings suggest potential therapeutic interventions, particularly highlighting PPAR-α agonists (fibrates) as promising candidates for treating advanced liver fibrosis in AATD patients, potentially opening up the treatment options for this proteotoxic disorder.
Publication
Nature
Authors
Florian A. Rosenberger, Sophia C. Mädler, Katrine Holtz Thorhauge, Sophia Steigerwald, Malin Fromme, Mikhail Lebedev, Caroline A. M. Weiss, Marc Oeller, Maria Wahle, Andreas Metousis, Maximilian Zwiebel, Niklas A. Schmacke, Sönke Detlefsen, Peter Boor, Ondřej Fabián, Soňa Fraňková, Aleksander Krag, Pavel Strnad, & Matthias Mann;
Title
Deep Visual Proteomics maps proteotoxicity in a genetic liver disease

