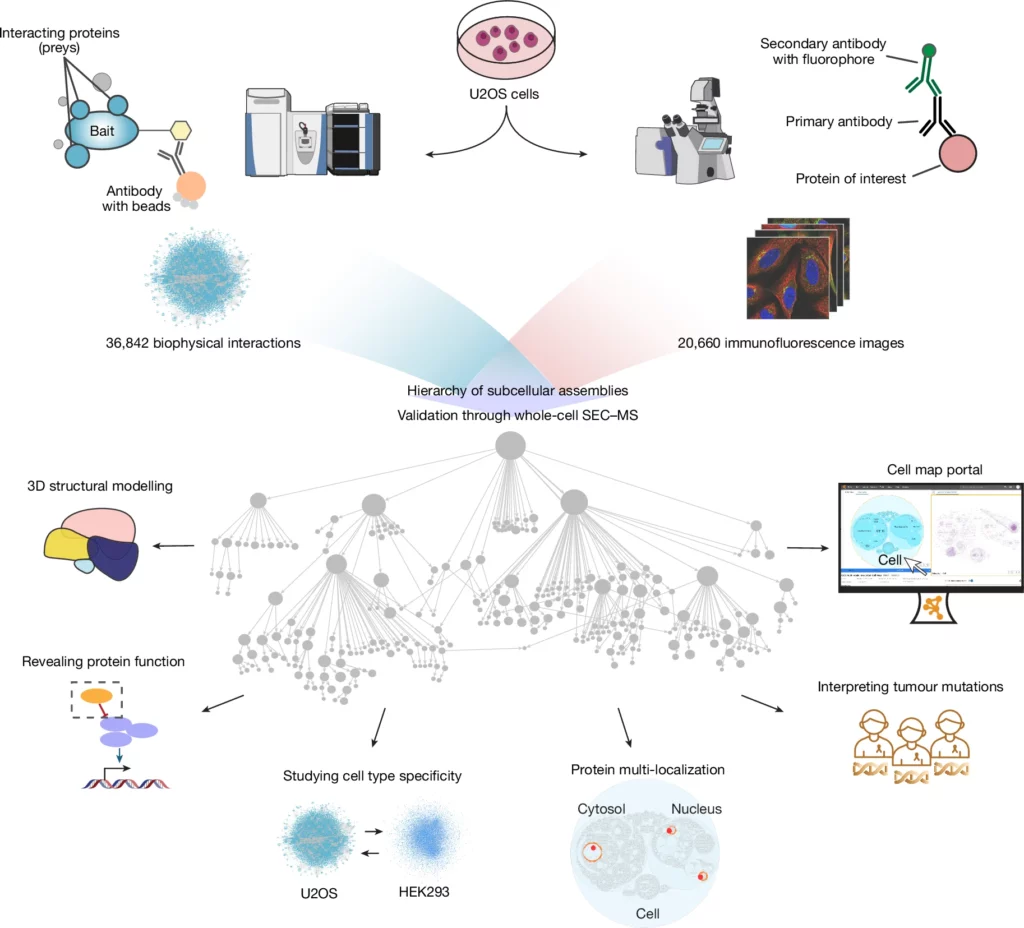
Study overview. From Schaffer et al., 2025. “Multimodal cell maps as a foundation for structural and functional genomics“, Nature (2025). https://doi.org/10.1038/s41586-025-08878-3. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Understanding the subcellular organisation of human cells is crucial for elucidating its relationship to biological function and disease. However, much of cellular architecture remains uncharted. To address this, researchers have employed a variety of technologies, including whole-cell electron microscopy, immunofluorescence staining and endogenous fluorescent tagging coupled with confocal microscopy, and mass spectrometry-based proteomics to investigate subcellular organization across multiple spatial scales. However, these approaches are typically applied independently, limiting the ability to achieve a comprehensive understanding of biological structures.
To overcome this challenge, Schaffer et al. created a comprehensive reference map of human protein subcellular organisation by integrating multiple proteomics data modalities. Using a machine learning approach, they combined confocal imaging data with affinity purification–mass spectrometry (AP-MS) interaction profiles for over 5,100 proteins in U2OS cells. This integrative strategy enabled the identification of 275 distinct molecular assemblies spanning sizes from 10⁻⁸ to 10⁻⁵ meters.
As orthogonal validation, the team employed size-exclusion chromatography–mass spectrometry (SEC-MS) to analyze cellular extracts from U2OS cells. 40 protein fractions were collected using a 1290 Series semi-preparative HPLC system. After tryptic digestion and peptide purification, each fraction was subjected to LC-MS analysis using a nanoElute UHPLC system coupled to a timsTOF Pro2 mass spectrometer. Peptide separation was performed on an IonOpticks Aurora Ultimate CSI 25×75 C18 UHPLC column. Analysis of the 40 individual fractions led to the identification of 5,509 proteins with high reproducibility across biological replicates. The SEC-MS results showed strong agreement with the multiscale cell map, confirming 89 assemblies—specifically, 43% of those containing more than 5 proteins and 61% of those containing more than 15 proteins.
This multimodal cell map demonstrates great promise as a foundational resource for broad applications across biology. It has enabled the structural determination of 111 heterodimeric protein complexes and provided new insights into the expanded architecture of the Rag–Ragulator assembly. Additionally, the map has functionally annotated 975 previously uncharacterized proteins, including newly identified roles for C18orf21 in RNA processing and DPP9 in interferon signaling, while also revealing multi-localized and cell-type-specific protein assemblies. Notably, the map has identified 21 recurrently mutated protein assemblies and implicated 102 newly validated cancer-associated proteins, providing critical insights into the genomic basis of pediatric cancers.
Publication
Nature
Authors
Leah V. Schaffer, Mengzhou Hu, Gege Qian, Kyung-Mee Moon, Abantika Pal, Neelesh Soni, Andrew P. Latham, Laura Pontano Vaites, Dorothy Tsai, Nicole M. Mattson, Katherine Licon, Robin Bachelder, Anthony Cesnik, Ishan Gaur, Trang Le, William Leineweber, Aji Palar, Ernst Pulido, Yue Qin, Xiaoyu Zhao, Christopher Churas, Joanna Lenkiewicz, Jing Chen, Keiichiro Ono, Dexter Pratt, Peter Zage, Ignacia Echeverria, Andrej Sali, J. Wade Harper, Steven P. Gygi, Leonard J. Foster, Edward L. Huttlin, Emma Lundberg & Trey Ideker
Title
Multimodal cell maps as a foundation for structural and functional genomics

