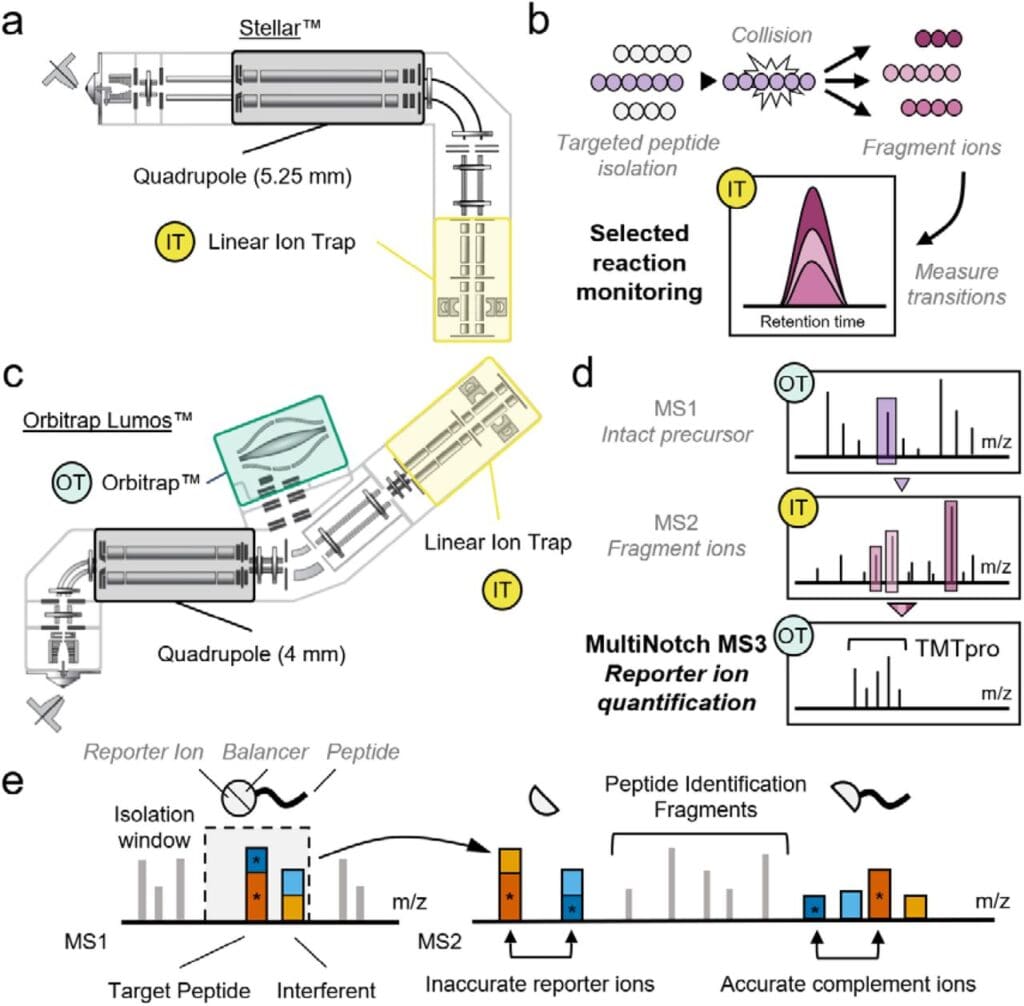
Comparison of Orbitrap Lumos & Stellar & their primary intended usage. From Cruz et al., 2025. “Expanding Targeted Instrumentation for Discovery Applications: Complement Reporter Ion Quantification with a Quadrupole-Ion Trap Instrument“, bioRxiv 2025.04.18.649483; doi: https://doi.org/10.1101/2025.04.18.649483. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
In proteomics research, multiplexed quantification methods using isobaric mass tags like TMTpro have traditionally required high-end mass spectrometers, limiting widespread adoption due to high costs. A significant knowledge gap existed in adapting these powerful techniques to more affordable instrumentation without compromising data quality.
Cruz et al. introduce iTMTproC (ion trap TMTproC), an adaptation of complement reporter ion quantification for quadrupole-ion trap instruments, enabling nine-channel multiplexed peptide quantification on cost-effective platforms. The researchers optimised various parameters including automatic gain control, ion injection time, and quadrupole isolation windows to achieve robust and accurate quantification.
The experimental approach involved benchmarking iTMTproC against established methods using both defined standards and complex biological samples. Analyses were performed on either a Stellar, Orbitrap Lumos, or Ascend mass spectrometer coupled to a Vanquish Neo UHPLC system or a Proxeon 1200 HPLC. Peptides were separated, across all instruments, with an IonOpticks Aurora Ultimate 25 cm × 75 μm C18 UHPLC column.
This quantitative study demonstrated that Stellar™ iTMTproC achieves comparable performance to Orbitrap Lumos™ MultiNotch MS3, an established method, on high-end instruments, quantifying a similar number of unique peptides while achieving 30% more protein identifications in single-shot analyses of a yeast–human standard mixture and 5% more proteins in fractionated complex biological samples, though with slightly lower quantification accuracy.
To assess the practical applicability of the method, the researchers applied it to investigate Xenopus laevis embryonic development and found that the protein dynamics measured by iTMTproC closely matched those obtained with MultiNotch MS3. The median cosine distance between temporal protein profiles was only 1.4 × 10⁻³, and it also demonstrated a high degree of consistency in the global protein expression trends between the two methods.
This advancement significantly enhances accessibility and promotes the democratisation of advanced quantitative proteomics, enabling a broader range of laboratories, without access to high-end mass spectrometry, to perform multiplexed shotgun proteomics using more affordable mass spectrometers.
Publication
bioRxiv
Authors
Edward R. Cruz, Alexander N.T. Johnson, Vyas Pujari, Thao Nguyen, Michael Stadlmeier, Jessica Wang, Cristina Jacob, Graeme C. McAlister, Philip M. Remes, & Martin Wühr;
Title
Advancing Multiplexed Proteomics on Cost-Effective Instrumentation

