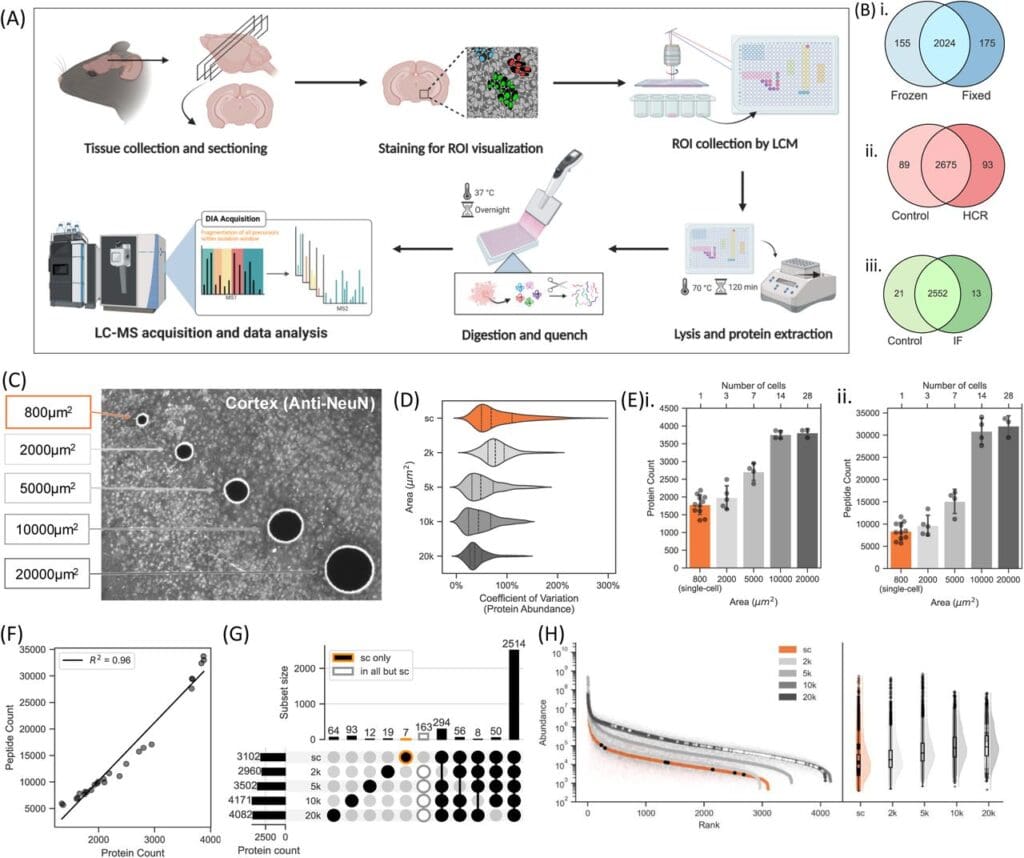
Design and optimisation of molecularly-guided spatial single-cell proteomics for CNS tissue. From Dutta et al., 2025. “Molecularly-guided spatial proteomics captures single-cell identity and heterogeneity of the nervous system“, bioRxiv 2025.02.10.637505; doi: https://doi.org/10.1101/2025.02.10.637505. Licensed under the terms of the Creative Commons CC-BY 4.0 license.
Understanding the precise molecular identity of spatially defined microenvironments has long challenged researchers in neurobiology and proteomics. The central nervous system presents a particularly unique obstacle: a complex interplay of highly specialised cells existing in tightly regulated and often disease-prone microenvironments. While transcriptomics has illuminated some of this complexity, resolving proteins with spatial fidelity at single-cell resolution could add a new perspective to further untangle this complexity.
Dutta et al.’s new method, LCM-ScP (Laser Capture Microdissection–Single-cell Proteomics), marries immunostaining-guided LCM with ultra-sensitive LC-MS to generate spatially resolved proteomic profiles of individual cells in both the central and peripheral nervous systems. The researchers used an IonOpticks Aurora Ultimate 25×75 C18 UHPLC column paired with a Vanquish Neo and an Orbitrap Exploris 480 with a Nanospray Flex ion source.
Using LCM-ScP, the researchers identified more than 2,000 proteins per neuron. They were able to uncover proteomic differences between closely related cell types in-situ, including elusive structures like ganglia and nerve bundles. The upregulation of specific proteins in defined neuronal subtypes – signals that would likely be obscured in bulk or transcriptomic analyses – highlights the sensitivity of the method and the strength of its validation.
This method offers a new lens to examine pathophysiology at the level of individual cells in dense microenvironments. It opens the door to biomarker discovery, subtype classification, and therapeutic targeting in previously inaccessible regions of the nervous system.
While challenges such as sample variability and potential contamination remain, this preprint serves as a proof-of-concept, demonstrating that transformational spatial single-cell proteomics is becoming increasingly accessible.
Authors
Sayan Dutta, Marion Pang, Gerard M. Coughlin, Sirisha Gudavalli, Michael L. Roukes, Tsui-Fen Chou, & Viviana Gradinaru;
Title



