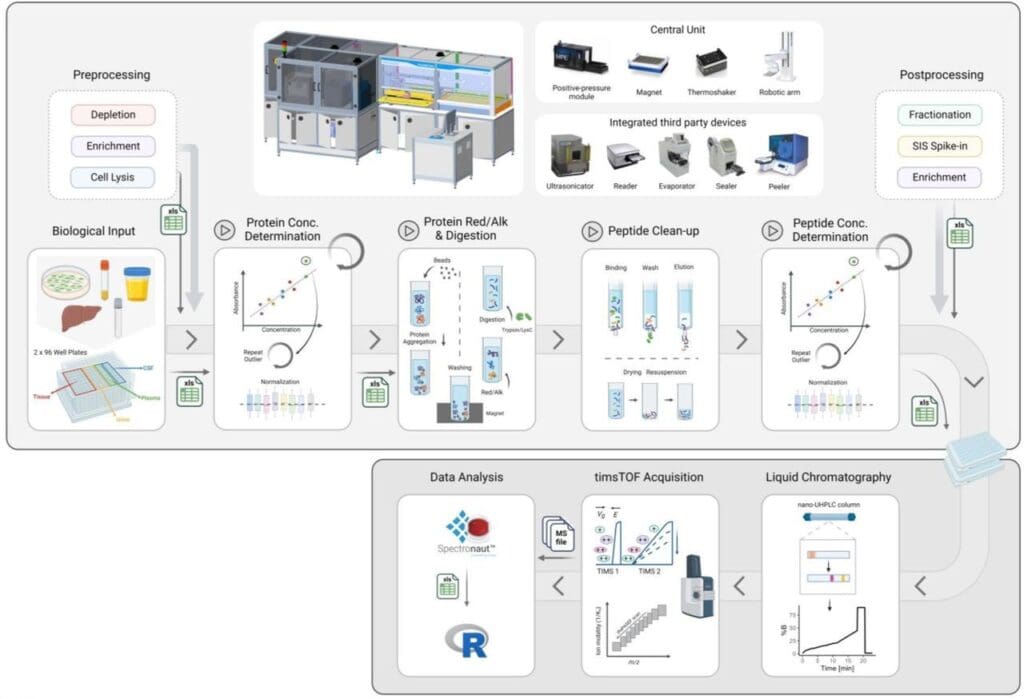
Schematic overview of the automated proteomics sample preparation platform coupled to short-gradient LC-MS acquisition. From Schär et al., 2025. “A flexible end-to-end automated sample preparation workflow enables reproducible large-scale bottom-up proteomics“, bioRxiv 2025.05.23.655690; doi: https://doi.org/10.1101/2025.05.23.655690. Licensed under the terms of the Creative Commons CC-BY-ND 4.0 license.
Bottom-up proteomics has emerged as a powerful tool for understanding disease mechanisms, but sample preparation remains a significant bottleneck limiting high-throughput applications. Traditional manual or semi-automated workflows are labor-intensive, prone to variability, and insufficient to meet the demands of large-scale clinical studies requiring the processing of hundreds of samples with consistent reproducibility.
To address these limitations, Schär, Räss et al. developed an automated, end-to-end sample preparation platform designed to process up to 200 samples per day with minimal hands-on time. The modular system integrates pre-processing, main sample processing, and post-processing blocks, accommodating diverse sample types including cultured cells, biofluids, and tissues from mid- to bulk-input levels. The platform utilises protein aggregation capture (PAC) protocol with automated liquid handling, temperature control, and error recovery systems.
Mass spectrometry analyses were performed using multiple instrument configurations tailored to specific acquisition modes. FAIMS-DIA experiments employed an EASY-nLC 1200 system coupled to an Orbitrap Exploris 480 mass spectrometer equipped with a Nanospray Flex ion source. For dia-PASEF acquisitions, an EASY-nLC 1200 was connected to a timsTOF HT with a CaptiveSpray II source. Targeted measurements were conducted using a Vanquish Neo UHPLC system coupled to an Exploris 480 with a FAIMS Pro interface. Chromatographic separation was carried out using various reversed-phase columns, including in-house packed columns, IonOpticks Aurora Frontier™ CSI 60 cm×75 μm C18 UHPLC column, and Aurora Ultimate™ CSI 25 cm×75 μm C18 UHPLC column.
This quantitative study demonstrated superior performance compared to established methods, achieving a 3.3-fold increase in throughput alongside a 6-fold reduction in manual intervention. The automated workflow showed improved protein identifications, with coefficients of variation ranging from 4-10% across replicates, and maintained consistent performance over several weeks of operation. The platform was successfully applied to molecular degrader screening experiments, enabling the identification of target proteins with degradation efficiencies ranging from 35% to 95%. This advancement enables routine large-scale proteomic studies essential for drug development and clinical applications, potentially accelerating biomarker discovery and therapeutic development.
Publication
bioRxiv
Authors
Sandra Schär, Luca Räss, Liliana Malinovska, Simonas Savickas, Francesca Cavallo, Christopher Below, Marco Tognetti, Polina Shichkova, Benoit Gourdet, Gonzalo Robles, Leo Iu, Jakob Vowinkel, Yuehan Feng, Roland Hjerpe, Roland Bruderer, Lukas Reiter
Title

